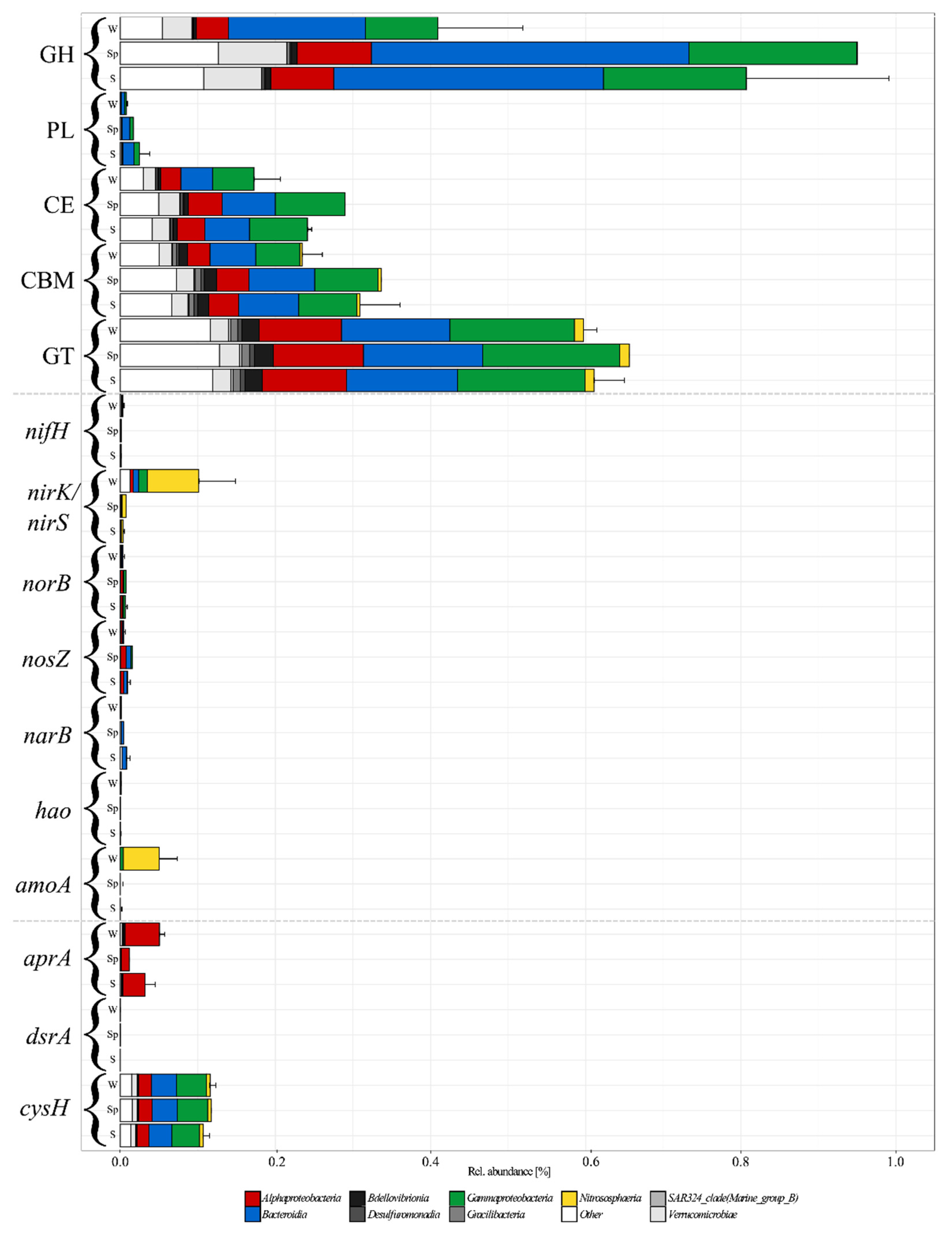
Microorganisms | Free Full-Text | A Winter-to-Summer Transition of Bacterial and Archaeal Communities in Arctic Sea Ice
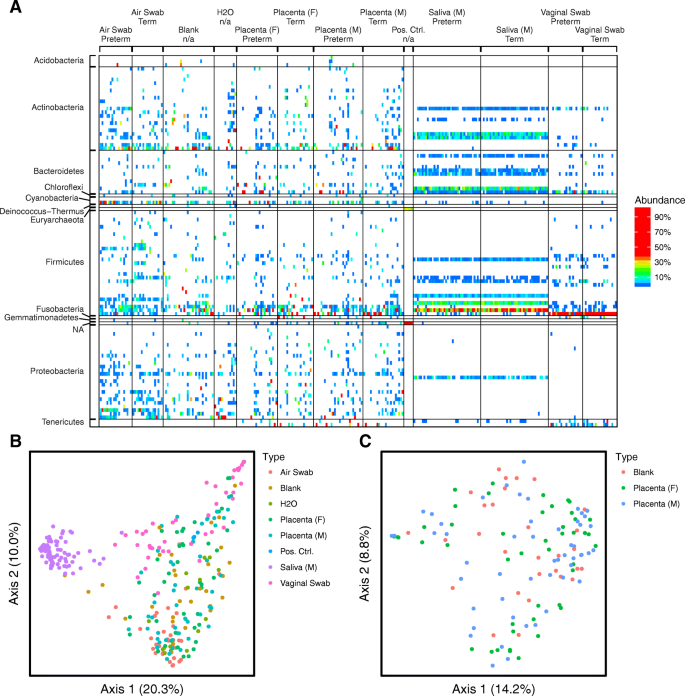
Lack of detection of a human placenta microbiome in samples from preterm and term deliveries | Microbiome | Full Text
![PDF] Simultaneous Quantification of Multiple Bacteria by the BactoChip Microarray Designed to Target Species-Specific Marker Genes | Semantic Scholar PDF] Simultaneous Quantification of Multiple Bacteria by the BactoChip Microarray Designed to Target Species-Specific Marker Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/2fe006dfd2eed4ad7bb10f2bc185631cbf45de5c/7-Figure3-1.png)
PDF] Simultaneous Quantification of Multiple Bacteria by the BactoChip Microarray Designed to Target Species-Specific Marker Genes | Semantic Scholar

A neural network‐based framework to understand the type 2 diabetes‐related alteration of the human gut microbiome - Guo - 2022 - iMeta - Wiley Online Library

Environmental Controls on Soil Microbial Communities in a Seasonally Dry Tropical Forest | Applied and Environmental Microbiology
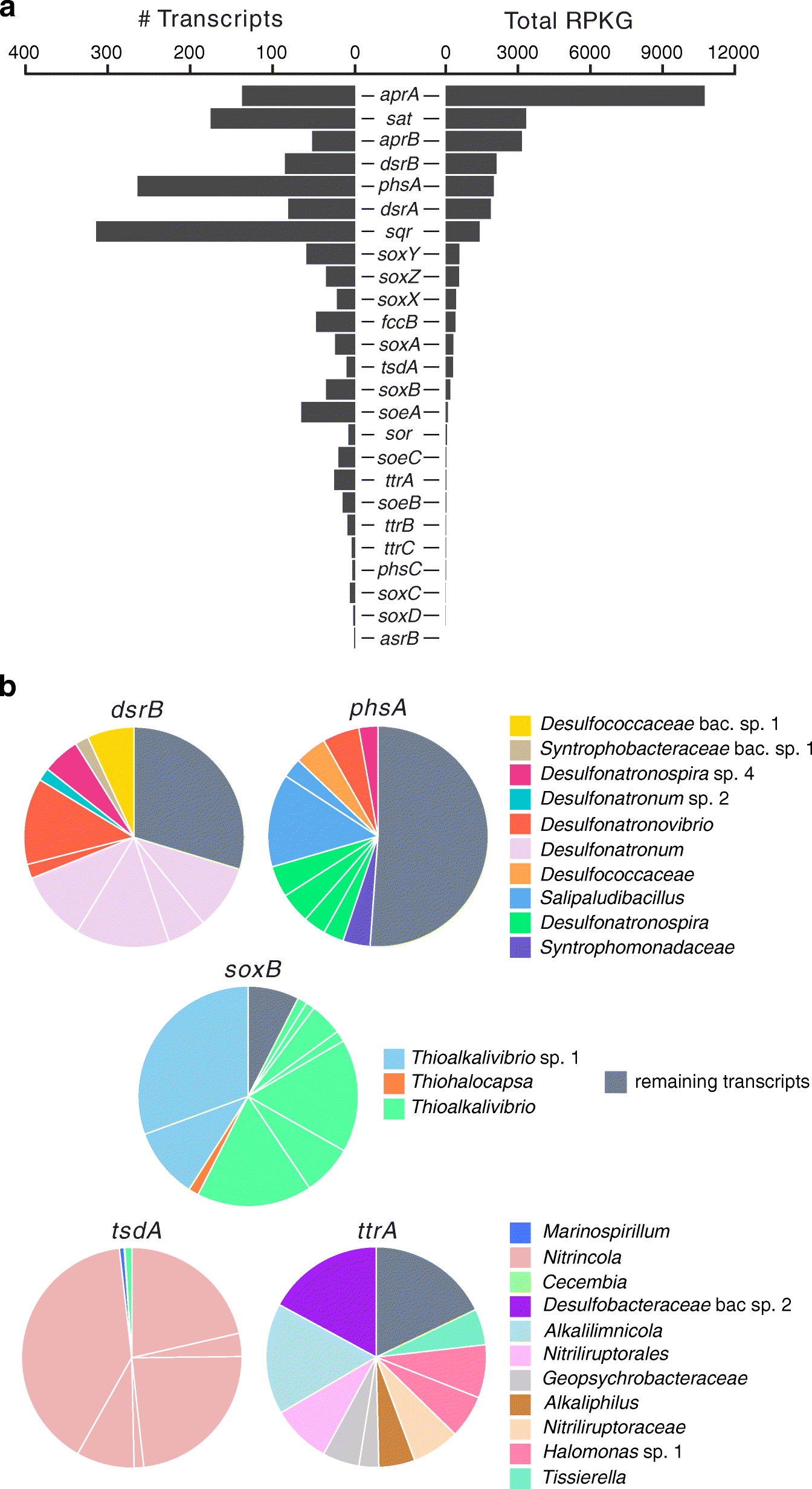
Metagenomes and metatranscriptomes shed new light on the microbial-mediated sulfur cycle in a Siberian soda lake | BMC Biology | Full Text
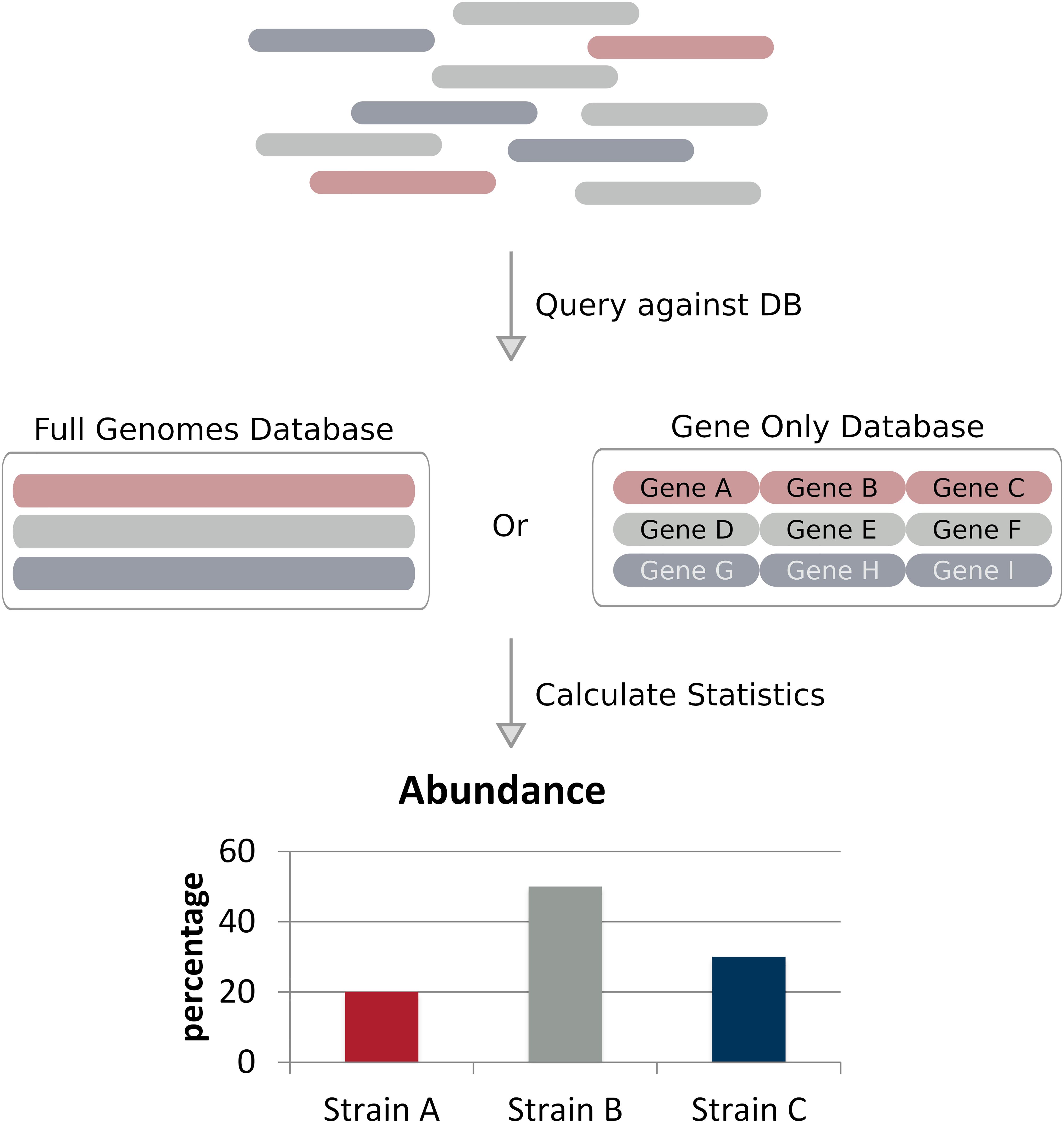
Frontiers | Computational Methods for Strain-Level Microbial Detection in Colony and Metagenome Sequencing Data

Virus-inclusive single-cell RNA sequencing reveals the molecular signature of progression to severe dengue | PNAS

Michael VanInsberghe on Twitter: "Cluster analysis and pseudotime ordering of these cells revealed not only the expected change in abundances of known cell-cycle marker genes, but also that the frequency of certain

𝙂𝙪𝙞𝙡𝙡𝙚𝙢 𝙎𝙖𝙡𝙖𝙯𝙖𝙧 on Twitter: "Very happy to publish this piece of work in which we discern the role of gene abundance vs. gene expression differences in structuring the global ocean microbial metatranscriptome.

Inferring gene regulation from stochastic transcriptional variation across single cells at steady state | PNAS
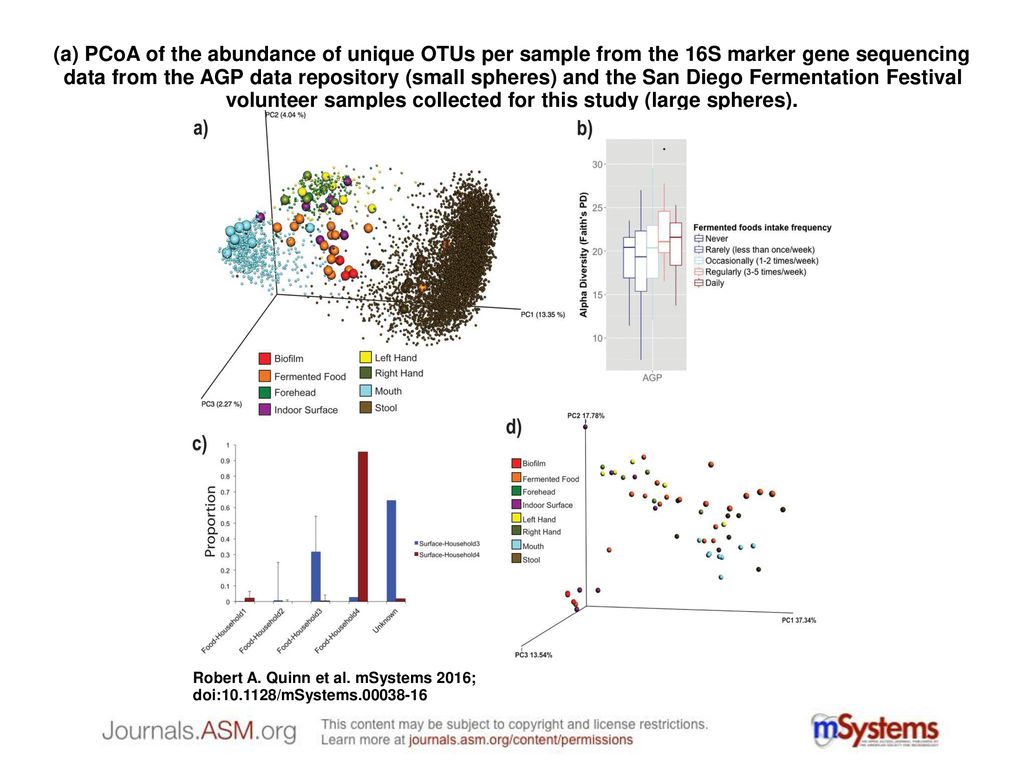
a) PCoA of the abundance of unique OTUs per sample from the 16S marker gene sequencing data from the AGP data repository (small spheres) and the San Diego. - ppt download




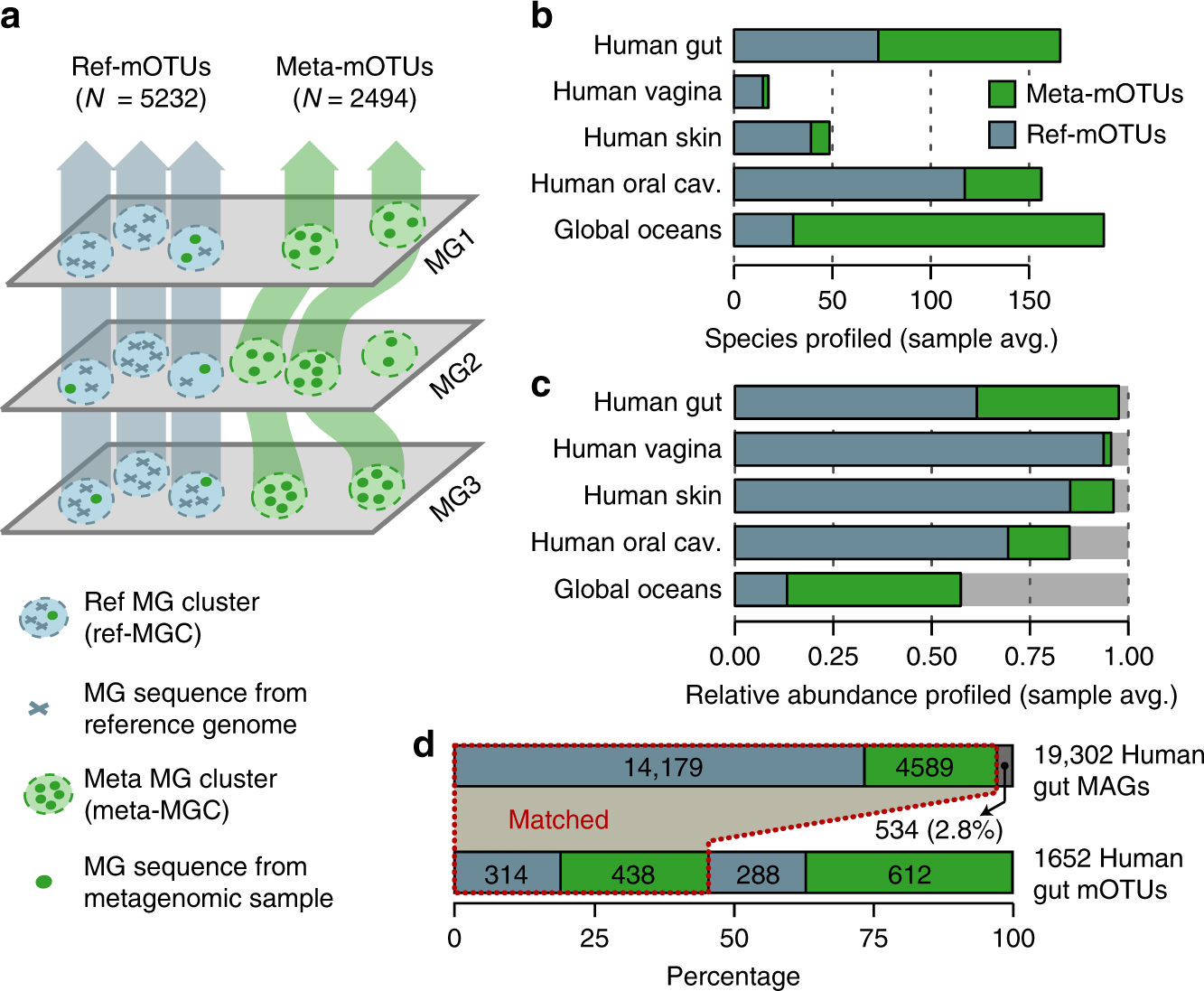
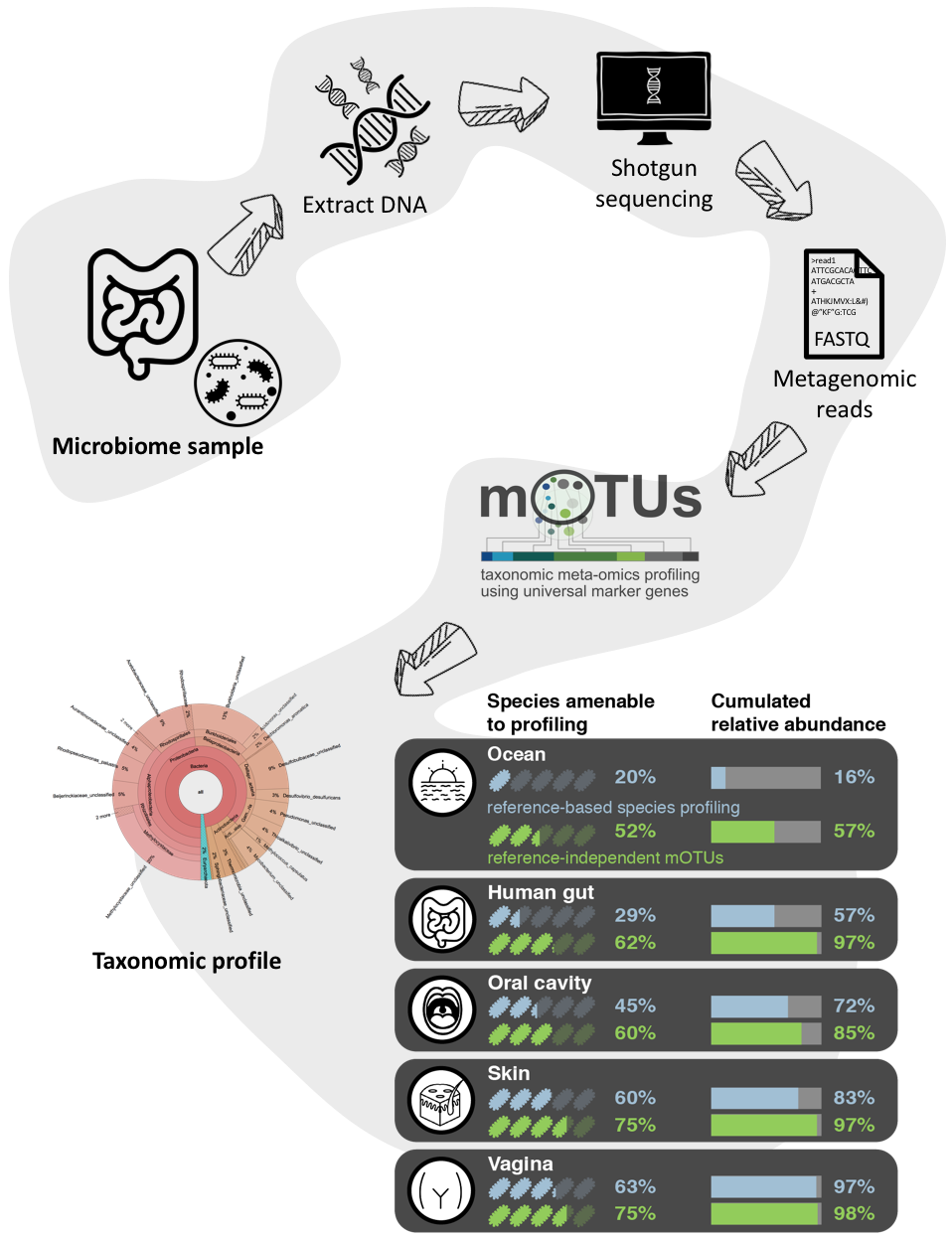

![Freshwater carbon and nutrient cycles revealed through reconstructed population genomes [PeerJ] Freshwater carbon and nutrient cycles revealed through reconstructed population genomes [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2018/6075/1/fig-1-full.png)

![PDF] TIPP2: metagenomic taxonomic profiling using phylogenetic markers | Semantic Scholar PDF] TIPP2: metagenomic taxonomic profiling using phylogenetic markers | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c471b9dd7bf0297cfe24555516a8770892c9072e/2-Figure1-1.png)

